A protocol for applying low-coverage whole-genome sequencing data
$ 11.99 · 4.5 (648) · In stock
STAR Protocols is an open access, peer-reviewed journal from Cell Press. We offer structured, transparent, accessible, and repeatable step-by-step experimental and computational protocols from all areas of life, health, earth and physical sciences.

Using low-coverage whole genome sequencing (genome skimming) to delineate three introgressed species of buffalofish (Ictiobus) - ScienceDirect
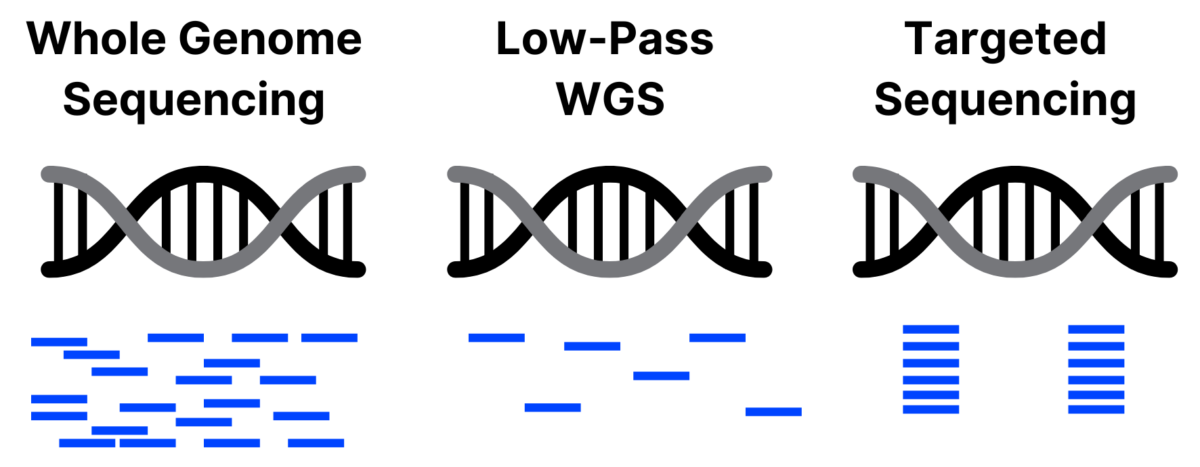
Low-pass sequencing and imputation for evaluating genetic variation - Gencove
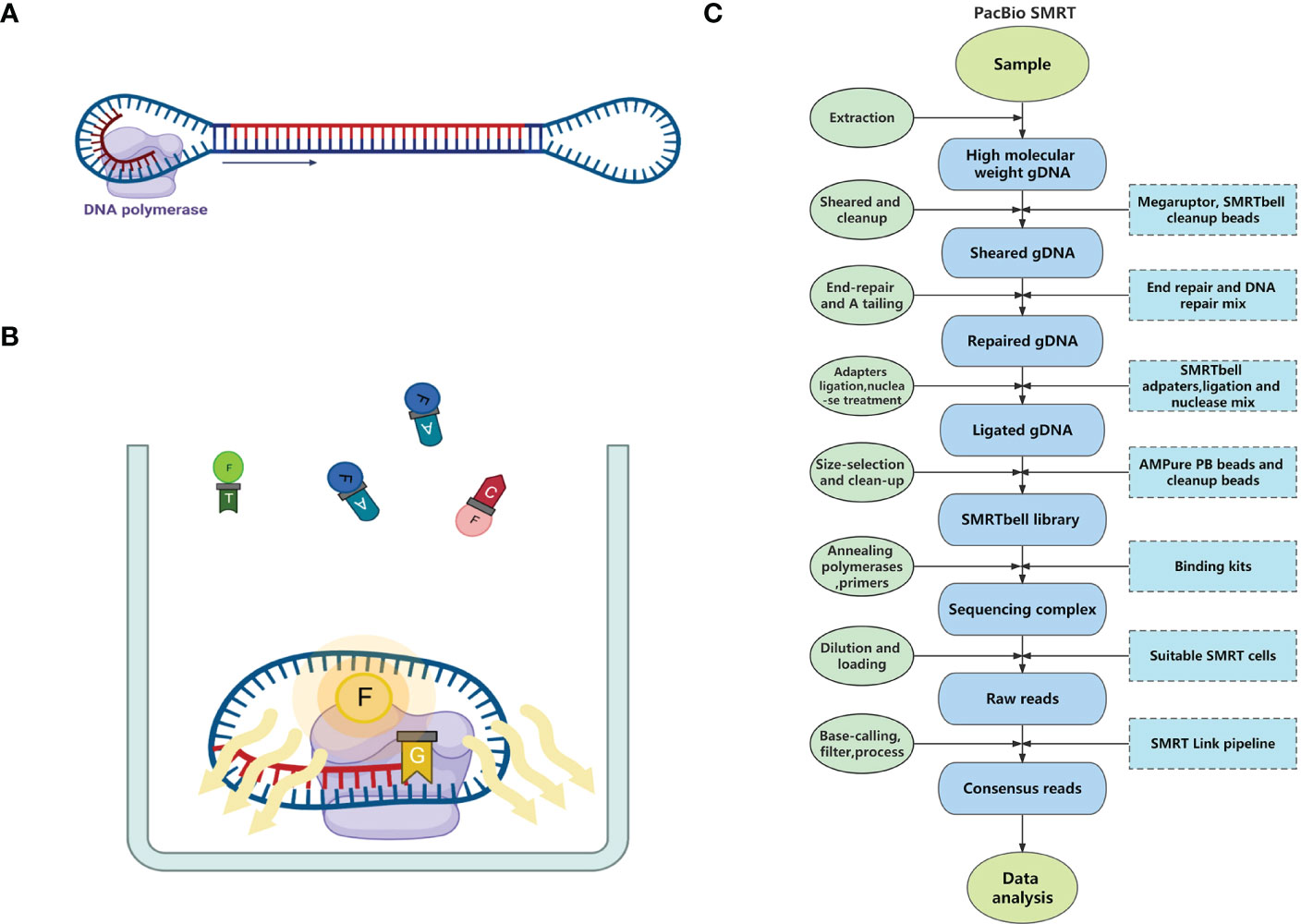
Frontiers Application of third-generation sequencing to herbal genomics
![]()
Shuhua Xu's research works Fudan University, Shanghai and other places

Copy number analysis by low coverage whole genome sequencing using ultra low-input DNA from formalin-fixed paraffin embedded tumor tissue, Genome Medicine

Hi-C (genomic analysis technique) - Wikipedia
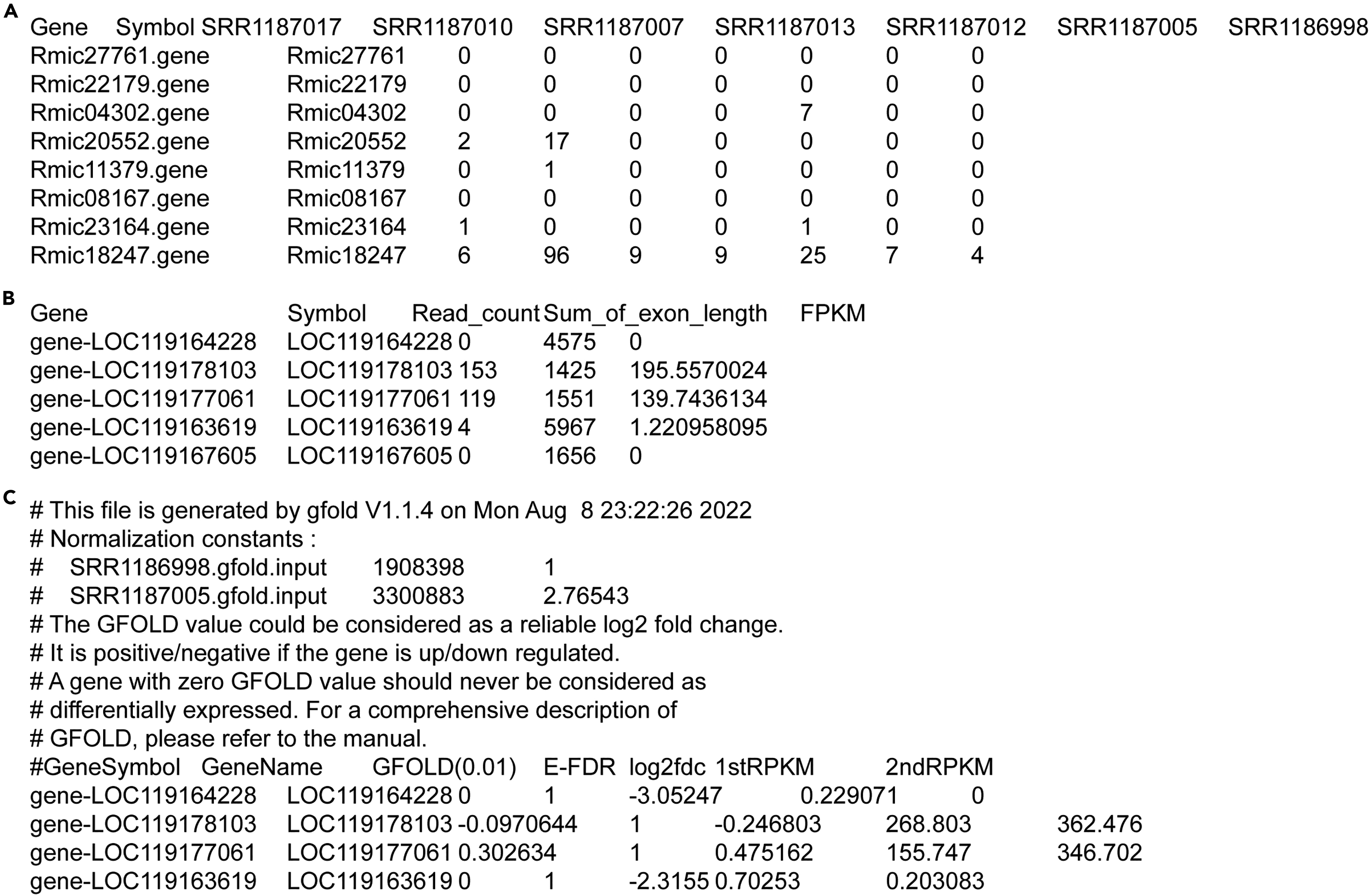

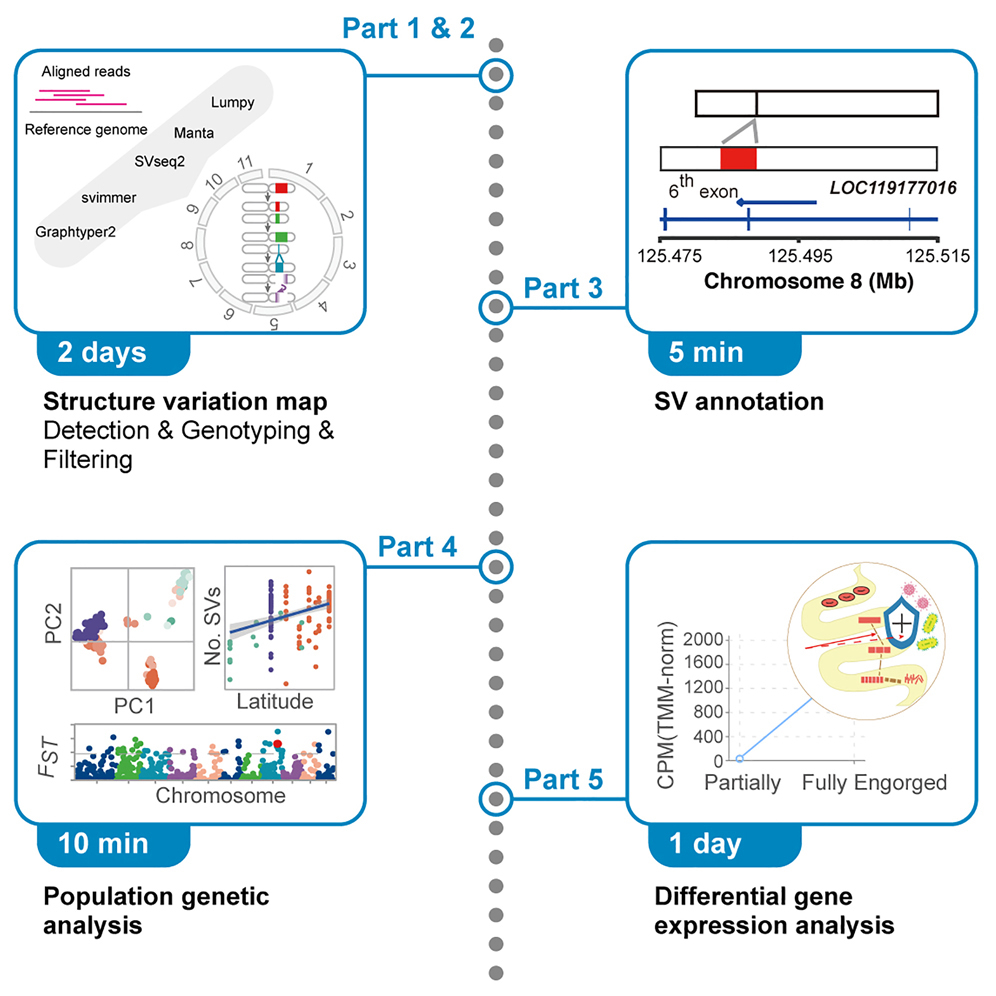
A protocol for applying low-coverage whole-genome sequencing data in structural variation studies

Yang GAO, PhD, Fudan University, Shanghai

Bo XIE, Chinese Academy of Sciences, Beijing, CAS, Shanghai Institutes for Biological Sciences
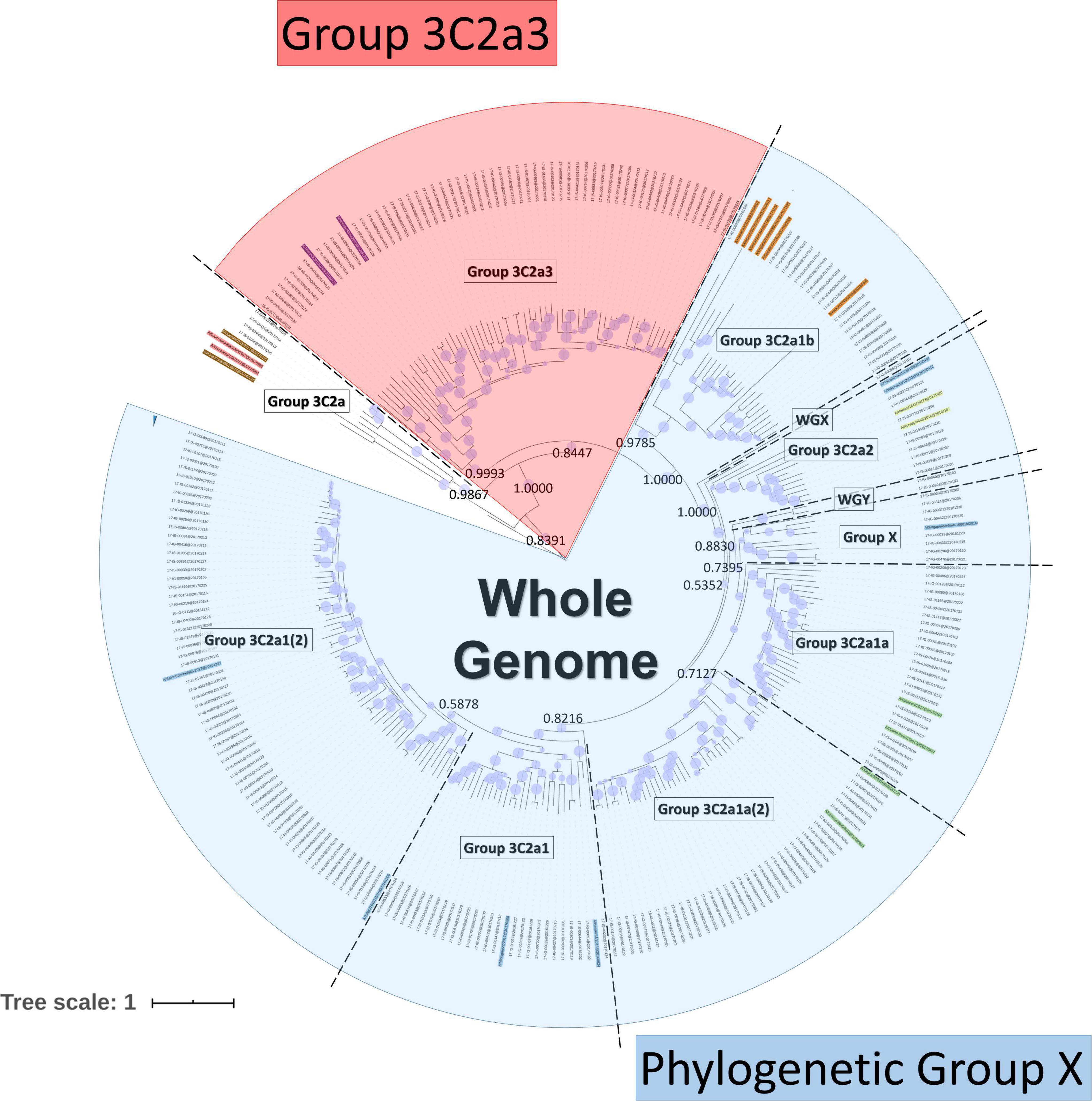
Frontiers Whole-Genome Sequence Approach and Phylogenomic Stratification Improve the Association Analysis of Mutations With Patient Data in Influenza Surveillance

PDF) PGG.SV: a whole-genome-sequencing-based structural variant resource and data analysis platform

9149 PDFs Review articles in RHIPICEPHALUS
Three Ways Gencove's Low-Pass Whole Genome Sequencing is Modernizing the Drug Discovery Process - Gencove
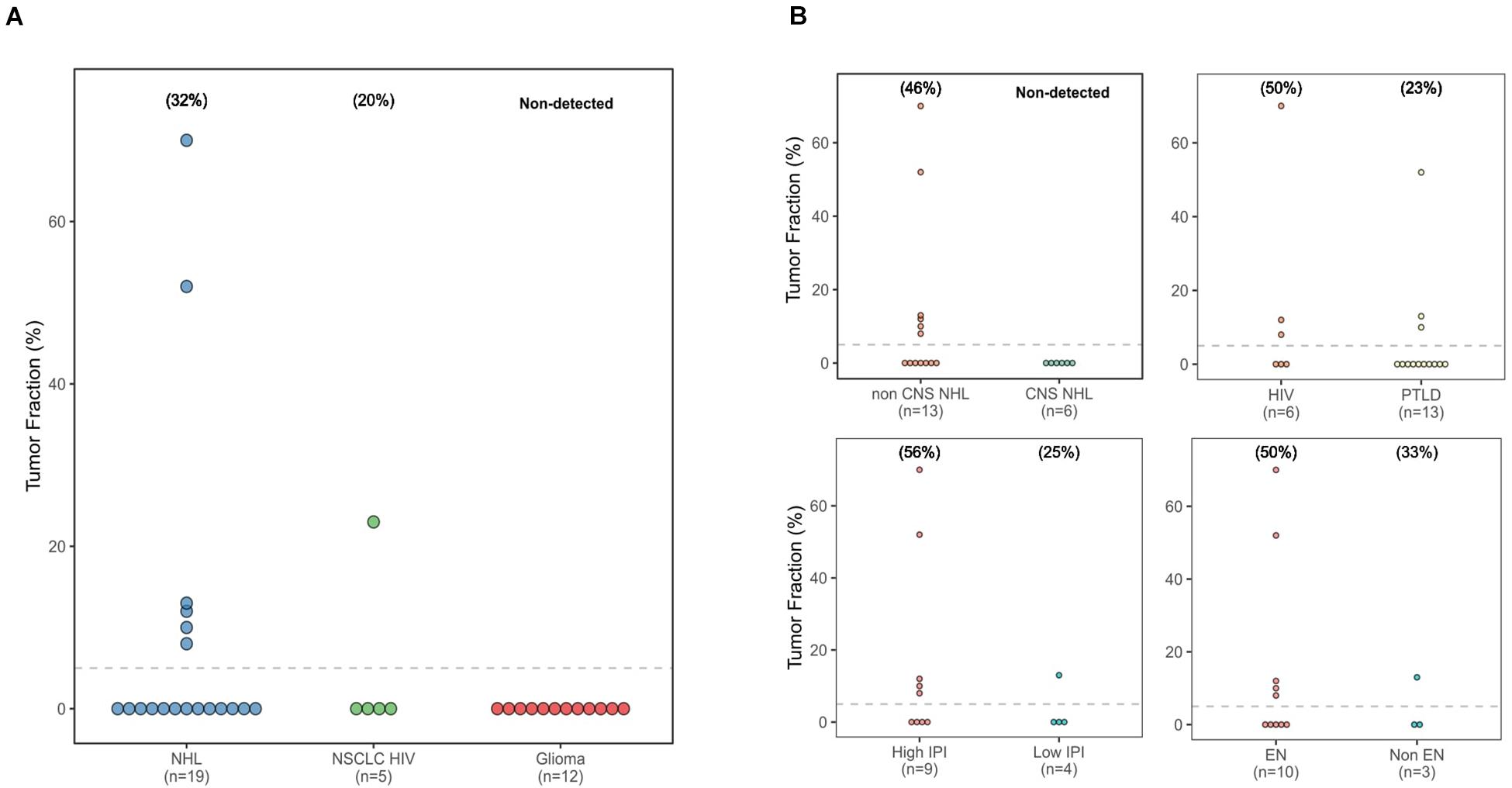
Frontiers Low-Coverage Whole Genome Sequencing of Cell-Free DNA From Immunosuppressed Cancer Patients Enables Tumor Fraction Determination and Reveals Relevant Copy Number Alterations